The currently circulating SARS-CoV-2 Omicron variant posed a big challenge for the ongoing pandemic prevention and control activities. Omicron variant showed resistance to neutralization by convalescent sera, vaccine-induced antibody and monoclonal antibodies including clinically approved monoclonal antibodies challenging the prevention and control activities.
Aggregation of more than 15 mutations in the receptor binding domain (RBD) of the Omicron spike alters its transmissibility, infectivity, neutralization escape, vaccine evasion, and reduced efficiency of diagnosis. RBD-induced immune evasion is common in Omicron owing to several factors. This variant demonstrated extreme evasion of most monoclonal antibodies tested. But it demonstrated insignificant evasion of neutralization a therapeutically approved monoclonal antibody (mAbs) sarbecovirus S309.
In a recent study published in Science, authors solved the X-ray crystal structure of a complex of the Omicron RBD with broadly neutralizing sarbecovirus mAb S309 (Figure 1). Mutations on the Omicron RBD (K417N, E484a and Q493R) are reported to abolish mAb binding through electrostatic contact rearrangements and steric hindrance. Besides, The Omicron RBD region comprising residues 366-375 (highlighted green in the Figure) is known to have deviated conformation compared to the Wuhan-Hu-1 RBD as it harbors S371L/S373P/S375F substitutions but without a significant effect on S309 neutralization. Overall, the results of this study entail that S309 binding mode remains unaltered by the Omicron mutations and the antibody effectively neutralized Omicron and other variants of concern.
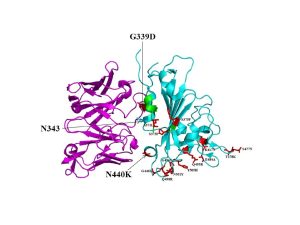
Figure 1: A cartoon representation of Omicron RBD (cyan) in complex with the therapeutically approved sarbecovirus monoclonal antibody S309 Fab (magenta) (PDB entry: 7TN0). G339D and N440K mutations found near or within the S309 antigenic recognition site are indicated in red sticks. S309 recognizes N343 glycan indicating non-significant alteration of S309 binding to Omicron RBD (McCallum, et al., 2022).
Journal article: McCallum M, et al., 2022. Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement. Science.
Summary by Hylemariam Mihiretie Mengist










