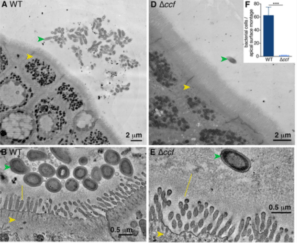
Donaldson et al., 2018. Figure 1 A,B,D &E. (A) Representative transmission electron microscopy (TEM) projection and (B) high-resolution tomogram of epithelial- associated wild-type B. fragilis in monocolonized mice. Ascending colons of mice harbored aggregates of B. fragilis (green arrow) under nonpathogenic conditions, that made tight associations with the glycocalyx (yellow line) overlying intestinal epithelial cells (IECs, yellow arrow). (D) Representative TEM projection image and (E) tomogram of epithelial-associated B. fragilis ∆ccf. The absence of the CCF system abrogated formation of bacterial aggregates and prevented intimate association with the glycocalyx. (n = 3 mice per group, about 1 mm epithelium scanned per mouse).
Immunoglobulin A (IgA) is the main antibody found in the gut. Studies about effects of IgA are largely in the context of infection or inflammatory disorders. These studies suggest that IgA may play a crucial role in shaping the intestinal microbiome, where IgA deficiency in mice has been associated with decreased microbiome diversity. The exact mechanisms by which IgA is involved in establishment and stability of the gut microbiome are poorly described.
Researchers aimed to determine what interactions occur between IgA and commensal bacterium that facilitate colonisation of gut mucosal surfaces. Bacteriodes fragilis (B.fragilis) is one of the major species of the gut microbiome. Researchers previously showed that wild-type (wt) B.fragilis deficient in commensal colonisisation factors (ΔccfB.fragilis) have reduced ability to colonise mucosal surfaces. In this research article showed that 7 of the 14 differentially expressed genes between mice colonised with wtB.fragilis and ΔccfB.fragilis encoded Ig variable chains. These transcriptomic differences were not associated with difference in immune responses with both wt– and ΔccfB.fragilis strains inducing similar levels of IgA. However, wtB.fragilis had significantly higher coating of IgA compared withΔccfB.fragilis and IgA antibodies induced by ΔccfB.fragilis were unable bind to wtB.fragilis. This shows that while some pathogenic bacteria may use capsular proteins for immune evasion, B.fragilis uses these proteins for colonisation in the gut. Additionally, mice colonised with ΔccfB.fragilis, were observed to have increased microbial diversity with reduced mucosal colonisation compared with wtB.fragilis.
In summary, this study shows that some microbial species such as B.fragilis from interaction with IgA, which results in increased colonisation. Additionally, the study also provides a mechanism as to why mice deficient in IgA or B cells have reduced Bacteriodes colonisation. Thus is non-pathogenic setting IgA enables colonisation of commensal bacteria.
Journal Article: Donaldson et al., 2018. Gut microbiota utilize immunoglobulin A for mucosal colonization. Science.
Article by Cheleka AM Mpande










