A new study has uncovered an unexpected molecular safeguard that helps the immune system tolerate dietary gluten, offering fresh insight into how Celiac disease develops (Figure 1). Researchers identified a long noncoding RNA called lnc13 as a key regulator of oral tolerance to gluten, revealing that loss of this molecule can drive immune changes that closely resemble human celiac disease.
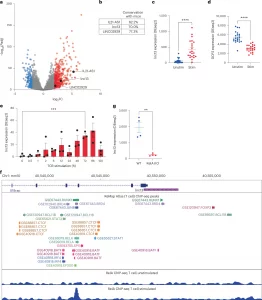
Figure 1: lnc13 is a candidate lncRNA regulator of oral tolerance to gluten.a, Volcano plot showing differentially expressed long-noncoding RNAs (lncRNAs) in gluten-specific CD4+ T cells from human small intestine activated by TCR for 3 h (data from GSE146441). n = 22 donors. lncRNAs upregulated (red) or downregulated (blue) are highlighted (abs|log2 fold change (FC)| > 2, adjusted P < 0.01). CeD-associated GWAS lncRNAs are shown in black. b, Cross-species sequence conservation (human-to-mouse alignment) of the three top candidate lncRNAs overlapping CeD-associated GWAS loci: IL21-AS1, LINC02929 and lnc13. c,d, Expression of lnc13 (C; P = 9.44537 × 10−7) (c) and its RNA-degrading enzyme DCP2 (D; P = 5.69524E-10) (d) in human gluten-specific CD4+ T cells at 0 and 3 h post-TCR stimulation (GSE146441). n = 22 donors. e, Time-course expression of human lnc13 in CD4+ T cells over 5 days of TCR activation (GSE96538), showing peak expression at days 3–4. n = 3 donors. P = 0.00078864. f, ChIP-seq analysis showing TF binding upstream of lnc13 in T cells. Middle: ReMap atlas of TF peaks; bottom: RELA ChIP-seq tracks in resting and 3-h-stimulated CD4+ T cells (GSE99319), indicating NF-κB recruitment. g, RNA expression of lnc13 in CD4+ T cells from WT and Rela-conditional knockout mice following 4 h of TCR stimulation (GSE239700), demonstrating NF-κB dependence. n = 4. P = 0.0017125. Differential expression (a,c–e,g) was computed using DESeq2 with adjusted P values that reflect Benjamini–Hochberg multiple comparisons correction. P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. In scatter-plots (c–e,g), lines represent mean ± s.e.m.
Celiac disease occurs when the immune system mounts an abnormal response to gluten, a protein found in wheat, barley, and rye. While it has long been known that genetic risk factors such as HLA-DQ2 and HLA-DQ8 are required for disease development, these genes alone do not fully explain why some individuals lose tolerance to gluten and develop chronic intestinal inflammation.
The new research points to a previously underappreciated layer of immune regulation involving long noncoding RNAs – RNA molecules that do not produce proteins but instead help control how genes are switched on and off.
The investigators screened for noncoding RNAs linked to immune activation and celiac disease–associated genetic variants, identifying lnc13 as a major candidate. To explore its function, they used HLA-DQ8 transgenic mice lacking lnc13 and examined how the animals responded to normal dietary gluten exposure.
Remarkably, simply feeding gluten to mice without lnc13 triggered immune and intestinal changes resembling hallmark features of celiac disease. The animals developed increased inflammatory IFN-γ–producing lymphocytes, elevated IL-12–expressing myeloid cells, expansion of cytotoxic intraepithelial immune cells, and crypt hyperplasia in the small intestine, all key signatures associated with loss of oral tolerance to gluten.
Mechanistically, the study showed that lnc13 acts as a molecular brake on inflammation by binding to regulatory DNA regions and limiting immune-cell responsiveness to pro-inflammatory signals. One particularly important pathway involved IL-15, a cytokine strongly implicated in celiac disease pathogenesis.
The researchers found that lnc13 restrains IL-15–driven differentiation of highly inflammatory CD8+ natural killer-like lymphokine-activated killer cells. Without lnc13, this inflammatory program becomes amplified, potentially pushing the immune system toward pathological gluten responses.
The findings highlight how noncoding elements of the genome can profoundly shape immune tolerance and intestinal homeostasis. They also suggest that defects in regulatory RNA pathways may contribute to the earliest stages of autoimmune disease development long before overt tissue damage appears.
Beyond celiac disease, the work adds to growing evidence that long noncoding RNAs are important regulators of immune balance and chronic inflammation. Understanding how molecules like lnc13 maintain tolerance could eventually open new therapeutic avenues aimed at restoring immune regulation rather than simply suppressing inflammation after disease onset.
Journal article: Yang-Fischer, R., et al. 2026. The long noncoding RNA lnc13 restrains inflammatory responses to maintain oral tolerance to gluten. Nature Immunology.
Summary by Stefan Botha










