The molecular arms race between bacteria and viruses has shaped some of the most important discoveries in modern biology. One of those discoveries, CRISPR, revolutionized gene editing by revealing how bacteria use specialized proteins to cut apart invading viral DNA.
But viruses are not passive targets. Over millions of years, they have evolved countermeasures known as anti-CRISPR proteins, molecular tools that disable bacterial defences and allow infection to proceed.
Now, researchers have uncovered an entirely new viral strategy, one that works not by blocking CRISPR after it is made, but by preventing the bacterial defence protein from ever existing in the first place (Figure 1).
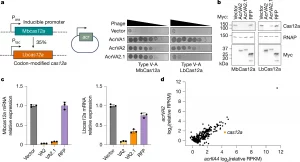
Figure 1: AcrVA2 specifically downregulates mRNA and protein of divergent cas12a orthologues.a, Schematic of strains engineered to express chromosomal Mbcas12a or codon-modified Lbcas12a from inducible promoters and acr genes from a plasmid (left). Phage plaque assay with tenfold serial dilutions of phage to assess CRISPR–Cas12a inhibition (right). Phage replication and subsequent host lysis causes clearances in the bacterial lawn. b, Western blot on bacterial lysates to assess the effect of Myc-tagged AcrVA2 or control proteins on Cas12a expression. RNAP, RNA polymerase (loading control). c, cas12a mRNA levels were quantified by qRT–PCR and normalized to rpoD. Expression is shown relative to empty vector control using the ΔΔCt method. Points represent technical triplicates (n = 3 technical replicates) from one representative experiment; mean ± s.d. is indicated. Experiments were independently repeated at least twice with similar results (a–c). d, Gene expression was quantified as reads per kilobase per million mapped reads (RPKM), normalized to the housekeeping gene rpoD, and log2 transformed. Each point represents one gene. Samples were processed simultaneously and the experiment was not repeated.
Published in Nature, the study reveals a remarkably precise mechanism in which a viral protein hijacks the bacterial ribosome, the cell’s protein factory, to selectively destroy a key CRISPR protein during its synthesis.
Most previously identified anti-CRISPR proteins work by binding directly to CRISPR-associated enzymes such as Cas proteins and preventing them from cutting viral DNA.
This newly identified viral protein, called AcrVA2, operates very differently.
Instead of targeting a finished CRISPR protein, AcrVA2 waits at the ribosome while proteins are being assembled. As soon as the ribosome begins producing the CRISPR protein Cas12, AcrVA2 recognizes the emerging amino acid chain and immediately intervenes.
The interaction effectively jams the ribosome mid-translation.
What makes the mechanism especially elegant is that the virus doesn’t directly destroy Cas12. Instead, it manipulates the bacterium’s own quality control systems.
Cells constantly monitor ribosomes for stalled or defective protein production. When AcrVA2 locks onto the growing Cas12 protein, the ribosome interprets the situation as a manufacturing error. This triggers a cleanup response that destroys:
- The partially synthesized Cas12 protein
- The Cas12 messenger RNA blueprint
As a result, the bacterial cell effectively eliminates its own antiviral defence machinery before it can be deployed.
One of the most striking aspects of the discovery is the specificity of the interaction.
AcrVA2 does not indiscriminately shut down protein synthesis. The viral protein appears to possess two distinct functional surfaces:
- One that anchors it to ribosomes
- Another that specifically recognizes newly forming Cas12 proteins
This allows the virus to selectively neutralize a critical immune defence without broadly damaging the host cell’s protein production machinery, an efficient strategy for maintaining a viable environment for viral replication.
According to the researchers, this may represent the first known example of one protein interrupting the production of another directly on the ribosome in such a targeted manner.
The finding expands how scientists think about both:
- Viral immune evasion
- Ribosome-associated quality control
It also highlights how sophisticated viral strategies can become in the evolutionary conflict between microbes and their pathogens.
Although the work focuses on bacterial immunity, the implications extend much further.
CRISPR systems underpin many modern gene-editing technologies, and understanding how anti-CRISPR proteins function could help researchers:
- Develop safer and more controllable CRISPR tools
- Create molecular “off-switches” for gene editing
- Better understand translation quality control mechanisms
The study also underscores a broader biological principle: viruses are master manipulators of cellular machinery, often revealing previously unknown aspects of cell biology in the process.
The discovery adds another chapter to the ongoing evolutionary battle between bacteria and viruses.
Bacteria evolved CRISPR to destroy viral genomes. Viruses evolved anti-CRISPR proteins to block CRISPR activity. Now, it appears some viruses have gone a step further, not merely inhibiting bacterial defenses but sabotaging their production at the source. In doing so, they reveal just how dynamic and inventive molecular evolution can be.
Journal article: Marino, N.D. et al. 2026. Translation-dependent degradation of cas12 mRNA triggered by an anti-CRISPR. Nature.
Summary by Stefan Botha










