Understanding T cells, how they activate, differentiate, and respond to disease, is central to modern immunology and the development of therapies like CAR-T. But the tools we rely on to study them haven’t kept pace. Traditional approaches such as flow cytometry are powerful, but come with trade-offs: they require fluorescent labelling, expensive instrumentation, and often destroy the cells in the process.
Now, researchers have developed a new method that could fundamentally change how we study immune cells, by imaging them live, label-free, and in real time (Figure 1).
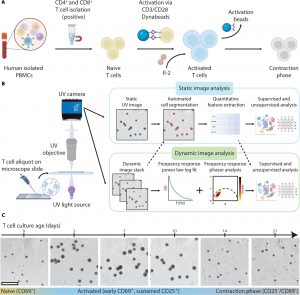
Figure 1: Deep-UV microscopy enables longitudinal imaging of live T cells. (A) Schematic of T cell isolation and activation protocol. (B) Sample preparation, imaging, and analysis workflow using deep-UV microscopy. (C) Sample deep-UV images of CD4+ T cells during a 21-d culture showing activation and subsequent contraction into post-activated cell phenotypes. Scale bar, 60 μm.
The team’s approach uses deep-ultraviolet (UV) microscopy at 255 nm, a wavelength strongly absorbed by nucleic acids. This enables high-contrast imaging of live T cells without any fluorescent dyes or destructive processing.
By combining this imaging with a custom-built neural network, the researchers could classify T cells into key functional states:
- Activated
- Dead
- Quiescent (naïve or contraction phase)
The performance was striking. Predictions closely matched traditional methods, with R² values above 0.97 for both viability and activation, effectively reproducing flow cytometry results without the drawbacks.
Identifying T cell subtypes, especially CD4-positive T cells versus CD8-positive T cells is a more complex problem. Static images alone weren’t enough.
To solve this, the researchers introduced dynamic deep-UV imaging, capturing rapid time-series data (500 frames at ~8 Hz).
These features, combined with UV absorption data, were fed into a second neural network. The result: ~90% accuracy in distinguishing CD4⁺ from CD8⁺ T cells, without any molecular markers.
One of the most interesting findings is that cellular metabolism itself becomes a detectable signal.
CD4⁺ T cells showed higher intracellular dynamic activity than CD8⁺ cells. This aligns with known biology:
- CD4⁺ cells rely more heavily on glycolysis and oxidative phosphorylation
- They contain more cytoplasmic mitochondria
The imaging data confirmed that these differences are localized to the cytoplasm, not the nucleus, directly linking the signal to metabolic machinery rather than genetic content.
This method offers a rare combination of advantages:
- Label-free: No dyes, no perturbation of the cells
- Non-destructive: Cells remain viable for downstream use
- High-throughput: Rapid imaging and analysis
- Mechanistically informative: Captures functional and metabolic states
For fields like immunotherapy, this is particularly valuable. In adoptive cell therapies (e.g., CAR-T), being able to monitor T cell activation and quality in real time without sacrificing cells could significantly improve manufacturing and clinical outcomes.
Journal article: Gorti, V, et al. 2026. Nondestructive, High-Resolution T Cell Characterization and Subtyping via Deep-UV Microscopy. BMEF.
Summary by Stefan Botha










